Main page
Introduction:
We’re a group of students: Chahd Maatallaoui, Yelyzaveta Larkina, Buddhima Senarathna, Laura Sommarberg, Onni Laukkala. We were assigned to do a Data Science Project with topic name “Cross-Species Transcriptomic Analysis of CUG-Repeat RNA to Identify Pathogenic Pathways in Myotonic Dystrophy”.
Our client for this project is Susana Garcia, who helped us with understanding biological aspect of our task and provided help during our work.
Our project is very scientific, so we decided to create this website to simplify and be able to explain what we have done. The research project is focused on working with data sets within molecular biology field, so we will provide explanation for difficult terms.
🧬 What is this project about? & Why this matters?
Our project focuses on myotonic dystrophy disease, which is characterized by muscle dysfunction and leads to disability and restriction of movements and causes potential dysfunction in the heart, brain, and other organs.
It can appear at any age, from birth to old age (0–70+ years) and symptoms usually first appear when individuals are in their 20s or 30s, but they can begin at any age and symptoms may not appear until 40 to 70 years of age.
There is no cure for this genetic disorder nowadays and our research work will contribute to finding what causes disruption in RNA-sequence work and hopefuly our results might be helpful to other researchers within this field as well.
And since it’s caused by toxic RNAs with expanded CUG repeats, but the molecular pathways remain unclear, we analyzed RNA-seq data to uncover which pathways are commonly disrupted, so we can see what causes this problem.
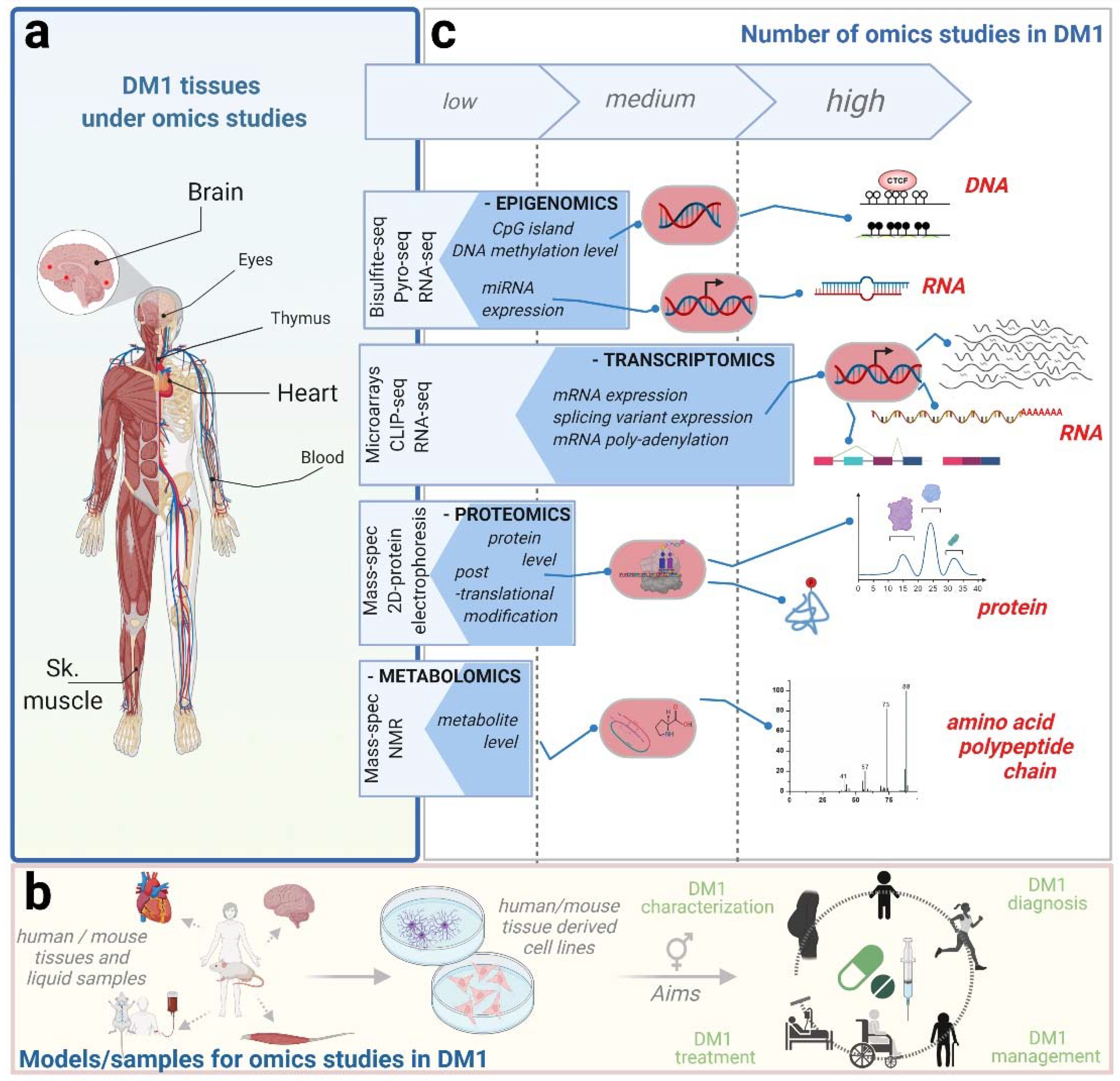
Researchers study these effects at multiple biological levels (DNA, RNA, protein). Credits for vizualization.
Explanation: This problem originates from mutations in the DNA. It is not the DNA itself, but the fact that it has been mutated, which leads—when it is read—to the production of an altered RNA compared to the wild-type form. While this change alone would not necessarily be problematic, the expanded-repeat RNA is toxic to the cell. .
Talking about biological aspect, we should mention that repetitive DNA sequences are very common in genomes, including the human genome. These repeat regions are naturally unstable and prone to errors when DNA is copied or read, which can lead to expansions or contractions in repeat length. Importantly, expansions of certain repeats are linked to an increasing number of neurodegenerative and neuromuscular disorders.
The CUG repeats that our group has focused on are one such example. In this case, expansions of CUG repeats cause Myotonic Dystrophy type 1 (DM1). In DM1—as in many repeat-associated disorders—the length of the expanded repeat strongly correlates with disease onset and severity: longer repeats (above the normal threshold) are associated with earlier onset and more severe symptoms.
For reference, repeat lengths in unaffected individuals typically range from about 5 to 37, whereas individuals with DM1 can have 50 to several thousand repeats.
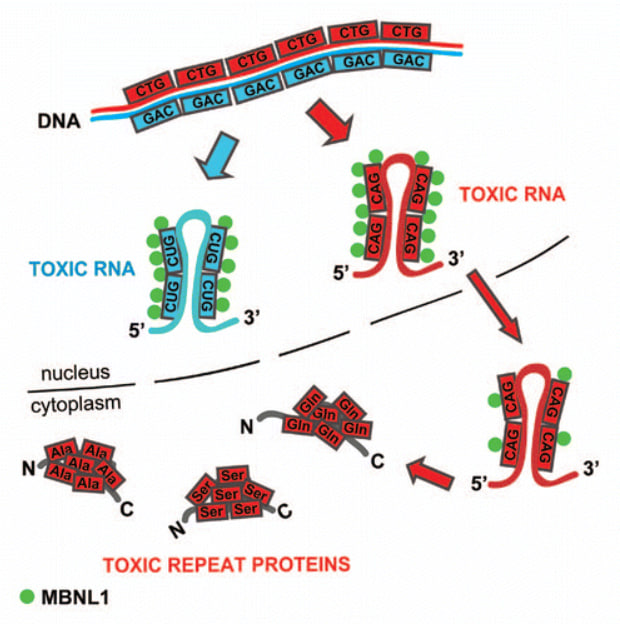
Example of expanded CTG repeats in DNA produce toxic RNA molecules with repeated CUG sequences. Credits for vizualization.
When the cell reads these expanded repeats, it produces RNA molecules that interfere with normal cellular processes. The RNA molecules produced from these expanded repeats disrupt multiple molecular and cellular processes, but the precise mechanisms by which they drive disease are still not fully understood. Emphasizing this uncertainty may also help convey why this research is important.
The main part of project includes working on worm data samples and then leaning towards working on human data.
🔗 Important links
Check how our working process went:
Check ui product made by our team member Chahd Maatallaoui, the main purpose of which is the reproducability of the result including multi organisms implementation. (will insert the link later when the product will be ready, but you can check the video of it in the meantime):
©️ Credits
Chahd Maatallaoui, Yelyzaveta Larkina, Buddhima Senarathna, Laura Sommarberg, Onni Laukkala - data scientists.
Susana Garcia - Biological consultant.
Chahd Maatallaoui - UI product developer.
Yelyzaveta Larkina - website developer.
Thanks everyone for help and support during this project!